-Search query
-Search result
Showing 1 - 50 of 53 items for (author: zhu & jk)

EMDB-35595: 
Cryo-EM structure of Cas12j-SF05-crRNA-dsDNA complex
Method: single particle / : Zhang X, Duan ZQ, Zhu JK

EMDB-28156: 
Cryo-EM structure of human DNMT3B homo-tetramer (form I)
Method: single particle / : Lu JW, Song JK

EMDB-28157: 
Cryo-EM structure of human DNMT3B homo-tetramer (form II)
Method: single particle / : Lu JW, Song JK

EMDB-28158: 
Cryo-EM structure of human DNMT3B homo-trimer
Method: single particle / : Lu JW, Song JK

EMDB-28159: 
Cryo-EM structure of human DNMT3B homo-hexamer
Method: single particle / : Lu JW, Song JK

EMDB-36579: 
A cryo-EM structure of the bovine chromogranin B dimer at a nominal resolution of ~0.35 nm
Method: single particle / : Jiang QX, Zhu MX, Yadav GP

EMDB-32490: 
SARS-CoV-2 spike glycoprotein trimer in closed state
Method: single particle / : Zhu Y, Tai L, Yin G, Sun F

EMDB-32491: 
SARS-CoV-2 spike glycoprotein trimer in closed state after treatment with Cathepsin L
Method: single particle / : Zhu Y, Tai L, Yin G, Sun F

EMDB-32492: 
SARS-CoV-2 spike glycoprotein trimer in Intermediate state
Method: single particle / : Zhu Y, Tai L, Yin G, Sun F

EMDB-32493: 
SARS-CoV-2 spike glycoprotein trimer in open state
Method: single particle / : Zhu Y, Tai L, Yin G, Sun F

EMDB-33832: 
CryoEM structure of Arabidopsis ROS1 in complex with TG mismatch dsDNA at 3.3 Angstroms resolution
Method: single particle / : Du X, Du J

EMDB-33835: 
CryoEM structure of Arabidopsis ROS1 in complex with 5mC-dsDNA at 3.1 Angstroms resolution
Method: single particle / : Du X, Du J

EMDB-33836: 
CryoEM structure of Arabidopsis ROS1 in complex with a covalent-linked reaction intermediate at 3.9 Angstroms resolution
Method: single particle / : Du X, Du J

EMDB-28558: 
SARS-CoV-2 BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment (local refinement of the RBD and S2X324)
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-28559: 
SARS-CoV-2 Omicron BA.1 spike ectodomain trimer in complex with the S2X324 neutralizing antibody Fab fragment
Method: single particle / : Park YJ, Seattle Structural Genomics Center for Infectious Disease (SSGCID), Veesler D

EMDB-30817: 
High Resolution Cryo-EM Structure of Cytochrome bo3 from E. Coli Reveals High Affinity Quinol Binding Site and Interactions of Protein with Lipids
Method: single particle / : Zhu JP, Zhang K, Gennis RB, Li J, Han L

EMDB-30818: 
High Resolution Cryo-EM Structure of Cytochrome bo3 from E. Coli Reveals High Affinity Quinol Binding Site and Interactions of Protein with Lipids
Method: single particle / : Zhu JP, Zhang K, Gennis RB, Li J, Han L

EMDB-30819: 
High Resolution Cryo-EM Structure of Cytochrome bo3 from E. Coli Reveals High Affinity Quinol Binding Site and Interactions of Protein with Lipids
Method: single particle / : Zhu JP, Zhang K, Gennis RB, Li J, Han L

EMDB-24265: 
E. coli cytochrome bo3 in MSP nanodisc
Method: single particle / : Vallese F, Clarke OB

EMDB-30471: 
2.55-Angstrom Cryo-EM structure of Cytochrome bo3 from Escherichia coli in Native Membrane
Method: single particle / : Li J, Han L, Gennis RB, Zhu JP, Zhang K

EMDB-30474: 
2.55-Angstrom Cryo-EM structure of Cytochrome bo3 from Escherichia coli in Native Membrane
Method: single particle / : Li J, Han L, Gennis RB, Zhu JP, Zhang K

EMDB-30475: 
Ubiquinol Binding Site of Cytochrome bo3 from Escherichia coli
Method: single particle / : Li J, Han L, Gennis RB, Zhu JP, Zhang K

PDB-7cub: 
2.55-Angstrom Cryo-EM structure of Cytochrome bo3 from Escherichia coli in Native Membrane
Method: single particle / : Li J, Han L, Gennis RB, Zhu JP, Zhang K

PDB-7cuq: 
2.55-Angstrom Cryo-EM structure of Cytochrome bo3 from Escherichia coli in Native Membrane
Method: single particle / : Li J, Han L, Gennis RB, Zhu JP, Zhang K

PDB-7cuw: 
Ubiquinol Binding Site of Cytochrome bo3 from Escherichia coli
Method: single particle / : Li J, Han L, Gennis RB, Zhu JP, Zhang K

EMDB-0667: 
Cryo-EM structure of the mammalian ATP synthase tetramer bound with inhibitory protein IF1
Method: single particle / : Gu JK, Zhang LX, Yi JB, Yang MJ

EMDB-0668: 
Cryo-EM structure of the mammalian E-state ATP synthase FO section
Method: single particle / : Gu JK, Zhang LX, Yi JB, Yang MJ

EMDB-0669: 
Cryo-EM structure of the mammalian E-state ATP synthase
Method: single particle / : Gu JK, Zhang LX, Yi JB, Yang MJ

EMDB-0670: 
Cryo-EM structure of the mammalian DP-state ATP synthase FO section
Method: single particle / : Gu JK, Zhang LX, Yi JB, Yang MJ

EMDB-0677: 
Cryo-EM structure of the mammalian DP-state ATP synthase
Method: single particle / : Gu JK, Zhang LX, Yi JB, Yang MJ

EMDB-6717: 
Contactin-2 reconstructed by Individual-Particle Electron Tomography
Method: electron tomography / : Lu Z, Liu JK, Ren G, Rudenko G

EMDB-9543: 
IPET 3D reconstruction of an single CNTNAP2-antibody complex: conformation #1
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9544: 
IPET 3D reconstruction of a single CNTNAP2-antibody complex: conformation #2
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9545: 
Single particle reconstruction of CNTNAP2: conformation #7
Method: single particle / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G
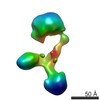
EMDB-9546: 
Single particle reconstruction of CNTNAP2: conformation #8
Method: single particle / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9547: 
IPET 3D reconstruction of a single CNTNAP2 - 1.8 nm nanogold complex
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9548: 
IPET 3D reconstruction of a single CNTNAP2 - 5 nm nanogold complex
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9549: 
IPET 3D reconstruction of a single CNTNAP2-C2 domain
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9550: 
Single particle reconstruction of CNTNAP2: conformation #6
Method: single particle / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9551: 
Single particle reconstruction of CNTNAP2: conformation #5
Method: single particle / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9552: 
Single particle reconstruction of CNTNAP2: conformation #4
Method: single particle / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9553: 
Single particle reconstruction of CNTNAP2: conformation #3
Method: single particle / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9554: 
Single particle reconstruction of CNTNAP2: conformation #2
Method: single particle / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9555: 
Single particle reconstruction of CNTNAP2: conformation #1
Method: single particle / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G
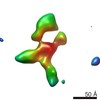
EMDB-9556: 
IPET 3D reconstruction of a single CNTNAP2: conformation #8
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G

EMDB-9557: 
IPET 3D reconstruction of a single CNTNAP2: conformation #7
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G
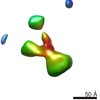
EMDB-9558: 
IPET 3D reconstruction of a single CNTNAP2: conformation #6
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G
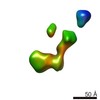
EMDB-9559: 
IPET 3D reconstruction of a single CNTNAP2: conformation #5
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G
EMDB-9560: 
IPET 3D reconstruction of a single CNTNAP2: conformation #4
Method: electron tomography / : Lu Z, Reddy MS, Liu JF, Kalichava A, Liu JK, Zhang L, Chen F, Wang Y, Holthauzen LM, Seshadrinathan S, Zhong XY, Ren G, Rudenko G
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search




 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
